News
How a Simple Genome Led to Phenotypic Complexity: Insights from Genomic and Epigenomic Studies in Aristolochia fimbriata
A research team led by Prof. JIAO Yuannian from the Institute of Botany of the Chinese Academy of Sciences (IBCAS) has created a high-resolution map of the centromeric architecture and three-dimensional (3D) epigenomic landscape of the magnoliid species Aristolochia fimbriata, offering new insights into how regulatory network rewiring drives the evolution of its specialized floral traits. The study was published inGenome Biology.
Aristolochia fimbriata is considered an exceptional model for studying plant evolution because, much like the basal angiosperm Amborella trichopoda, it has not undergone additional whole-genome duplication (WGD) events since the origin of extant flowering plants. Therefore, it could serve as a "living fossil" for inferring ancestral genetic features. Owing to its short life cycle, ease of cultivation, and relatively small genome, this species has been proposed as a promising model for early-diverging angiosperms. Notably, it exhibits highly specialized floral organs, unique coloration and scent, and an unusual "trap–imprison–release" pollination system, raising the question of how such complex traits evolved from a simple genome.
In this study, the researchers generated a telomere-to-telomere (T2T) genome assembly of A. fimbriata. A standout discovery was the identification of remarkably short, 34-bp centromeric satellite repeats (CEN34). These represent the shortest centromeric monomers reported among all investigated plant T2T genomes to date. The study also revealed that these centromeres are highly homogenized and almost entirely devoid of transposable elements (TEs), challenging traditional views on centromere complexity and evolution.
This research marks the first report of topologically associating domain-like (TAD-like) structures in the magnoliid clade. Using high-resolution Hi-C data, the team identified approximately 1,020 of these TADs, which help organize the genome into functional neighborhoods. These features contrast sharply with the largely TAD-less genome organization of Arabidopsis, demonstrating that prominent TAD structures can exist even in compact plant genomes. The 3D genome also led to key findings. Over 50% of accessible chromatin regions (ACRs) participate in distal chromatin loops that bypass neighboring genes to regulate distant targets. A significant correlation was found between gene looping and transcriptional activity, suggesting that the physical folding of DNA directly shapes gene expression.
To understand the genetic basis of A. fimbriata's elaborate flowers, the team analyzed the regulatory network of the B-class floral organ identity gene AfAP3. Compared to model plants like Arabidopsis, the downstream regulatory network of AfAP3 in A. fimbriata has significantly expanded to include pathways for anthocyanins, carotenoids, and terpenes. These findings indicate that despite having a simplified genome, A. fimbriata exhibits a more complex regulatory network, and that the rewiring of three-dimensional genome architecture and gene regulatory networks may represent an evolutionary strategy of "achieving complexity with simplicity."
This study not only establishes A. fimbriata as a powerful genetic model for the magnoliid lineage but also provides a foundational resource for comparative genomics across all flowering plants. The complete T2T genome and epigenetic maps will enable scientists to further explore how 3D chromatin reorganization and regulatory rewiring drive phenotypic innovation and ecological adaptation.

Flower of Aristolochia fimbriata
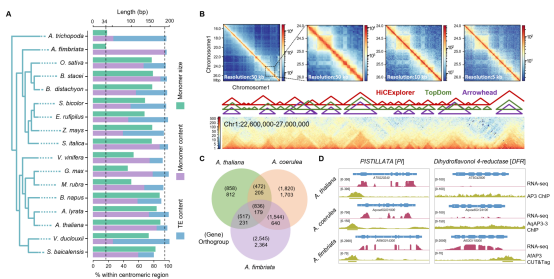
Evolution of centromeres, 3D genome architecture, and regulatory network expanding in Aristolochia fimbriata.




