Domesticated Asian rice (Oryza sativa L.), consisting of two subspecies (Japonica and Indica), is not only one of most important crops worldwide but also an excellent model of biological research. Despite extensive investigations, the origin/domestication of Asian rice has been controversial for almost one century. Of various opinions or arguments, two leading hypotheses (the single vs. multiple domestications) have been widely acknowledged but remain unsolved. In a study published in Nature Plants on August 7, a research team led by Prof. GE Song from the Institute of Botany of the Chinese Academy of Sciences (IBCAS) conducted extensive analyses on rice domestication history at the genome scale based on a dataset of 459 newly resequenced and 1119 publicly available genomes of wild and cultivated accessions.
By proposing a new strategy to test for alternative hypotheses, the researchers identified 993 selected genes that were shared by Japonica and Indica, suggesting that the domestication alleles of these genes originated only once in either Japonica or Indica. Importantly, researchers revealed that the domestication alleles of most selected genes (~80%) stemmed from wild rice O. rufipogon in China, but the domestication alleles of a substantial minority of selected genes (~20%) originated from wild rice O. nivara in South and Southeast Asia, demonstrating separate domestication events of Asian rice in different areas (Figure 1).
In addition, this study contributed genomic resources, including 422 newly resequenced wild accessions verified by phenotypic examination, which will greatly facilitate further studies of rice genetics, evolution and breeding. The extended dataset allowed the authors to uncover deep genetic structure and multiple distinct lineages within both wild and domesticated rice, which ensures a better phylogenetic framework to infer evolutionary history of domestication genes. Unexpectedly, the researchers found that some selective sweep regions in rice contained genes from different evolutionary origins, cautioning against previous practice of reconstructing a single phylogeny based on concatenating sweep regions in inference of domestication history.
"This study provide evidence that rice domestication began independently from divergent wild lineages in different areas, which was then completed with continuous selection and the exchange of beneficial alleles among different cultivar groups. This study also provides important implications for studies of other domesticated species, particularly for those with confounded evolutionary histories defined by widespread and continuous gene flow among cultivars and lines",said Prof. GE Song, the corresponding author.
The study was funded by the National Natural Science Foundation of China and the CAS Strategic Priority Research Program and Ministry of Science and Technology.
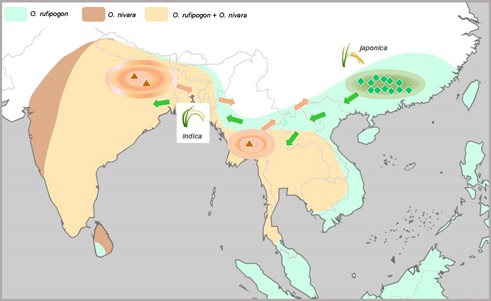
Figure 1. Hypothesized domestication centers of domesticated Asian rice. (Image by IBCAS) Geographic distributions of two ancestral species (O. rufipogon and O. nivara) were shaded in different colors. Origin and dispersal of the domesticated alleles shared by japonica and indica were depicted, with those in green diamond domesticated in japonica and those in brown triangle in indica.
Article Link: https://www.nature.com/articles/s41477-023-01476-z
Contact:
Prof. GE Song, gesong@ibcas.ac.cn
The Institute of Botany, Chinese Academy of Sciences
Domesticated Asian rice (Oryza sativa L.), consisting of two subspecies (Japonica and Indica), is not only one of most important crops worldwide but also an excellent model of biological research. Despite extensive investigations, the origin/domestication of Asian rice has been controversial for almost one century. Of various opinions or arguments, two leading hypotheses (the single vs. multiple domestications) have been widely acknowledged but remain unsolved. In a study published in Nature Plants on August 7, a research team led by Prof. GE Song from the Institute of Botany of the Chinese Academy of Sciences (IBCAS) conducted extensive analyses on rice domestication history at the genome scale based on a dataset of 459 newly resequenced and 1119 publicly available genomes of wild and cultivated accessions.
By proposing a new strategy to test for alternative hypotheses, the researchers identified 993 selected genes that were shared by Japonica and Indica, suggesting that the domestication alleles of these genes originated only once in either Japonica or Indica. Importantly, researchers revealed that the domestication alleles of most selected genes (~80%) stemmed from wild rice O. rufipogon in China, but the domestication alleles of a substantial minority of selected genes (~20%) originated from wild rice O. nivara in South and Southeast Asia, demonstrating separate domestication events of Asian rice in different areas (Figure 1).
In addition, this study contributed genomic resources, including 422 newly resequenced wild accessions verified by phenotypic examination, which will greatly facilitate further studies of rice genetics, evolution and breeding. The extended dataset allowed the authors to uncover deep genetic structure and multiple distinct lineages within both wild and domesticated rice, which ensures a better phylogenetic framework to infer evolutionary history of domestication genes. Unexpectedly, the researchers found that some selective sweep regions in rice contained genes from different evolutionary origins, cautioning against previous practice of reconstructing a single phylogeny based on concatenating sweep regions in inference of domestication history.
"This study provide evidence that rice domestication began independently from divergent wild lineages in different areas, which was then completed with continuous selection and the exchange of beneficial alleles among different cultivar groups. This study also provides important implications for studies of other domesticated species, particularly for those with confounded evolutionary histories defined by widespread and continuous gene flow among cultivars and lines",said Prof. GE Song, the corresponding author.
The study was funded by the National Natural Science Foundation of China and the CAS Strategic Priority Research Program and Ministry of Science and Technology.

Figure 1. Hypothesized domestication centers of domesticated Asian rice. (Image by IBCAS) Geographic distributions of two ancestral species (O. rufipogon and O. nivara) were shaded in different colors. Origin and dispersal of the domesticated alleles shared by japonica and indica were depicted, with those in green diamond domesticated in japonica and those in brown triangle in indica.
Article Link: https://www.nature.com/articles/s41477-023-01476-z
Contact:
Prof. GE Song, gesong@ibcas.ac.cn
The Institute of Botany, Chinese Academy of Sciences
